Project Proposal
Motivation
One of the biggest bottlenecks in machine learning pipelines is the amount of labeled data necessary to train a model. The problem setup is we have a large unlabeled corpus of documents and can ask an annotator to provide labels for the documents however there is an associated cost to annotate a document. The goal is to maximize model performance and minimize annotation cost. There have been various methods to reduce the amount of data needed to train machine learning models such as: transfer learning, active learning, and weak supervision to name a few. This project focuses on combining those 3. The goal being, as a user hand labels a set of instances, we generate a set of labeling functions that learn heuristics about the instances that have been labeled and apply it to the entire corpus, to get additional signal from unlabeled instances. The generic pipeline is described in figure 1.
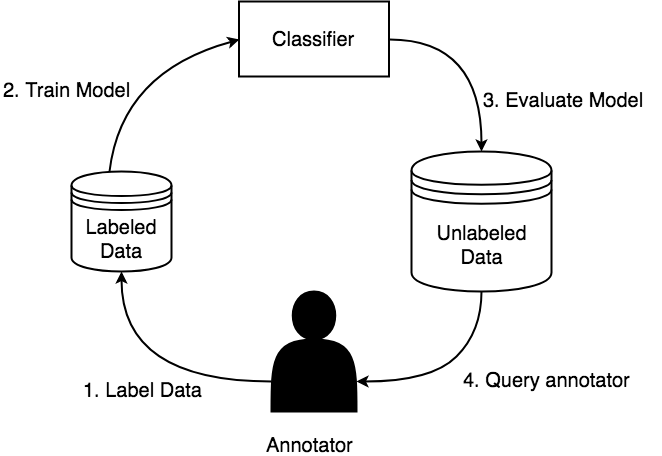
Figure 1: Description of a generic active learning workflow. (1) end user annotates data, (2) from the annotated data a model is trained, (3) the model is evaluated on the entire corpus, (4) a new query of data for the user to annotated is selected based on some heuristic, then process repeats
In particular, we take a look at binary sequence classification problems such as NER (named entity recognition). We call our approach DPD (data programming by demonstration) as this is an application of programming by demonstration in the language domain. We build on top of Snorkel [1 Ratner et al. 2017], a machine learning platform that introduces data programming where users write python functions to label their data, and train models to learn from it. This process is described in figure 2. We follow the traditional active learning pipeline (figure 1), where the training set is iteratively built up and a model is trained on it. During training we not only the use annotated data, but also the annotated instances to generate a set of labeling functions and apply to the unlabeled corpus to hopefully get more signal during training.
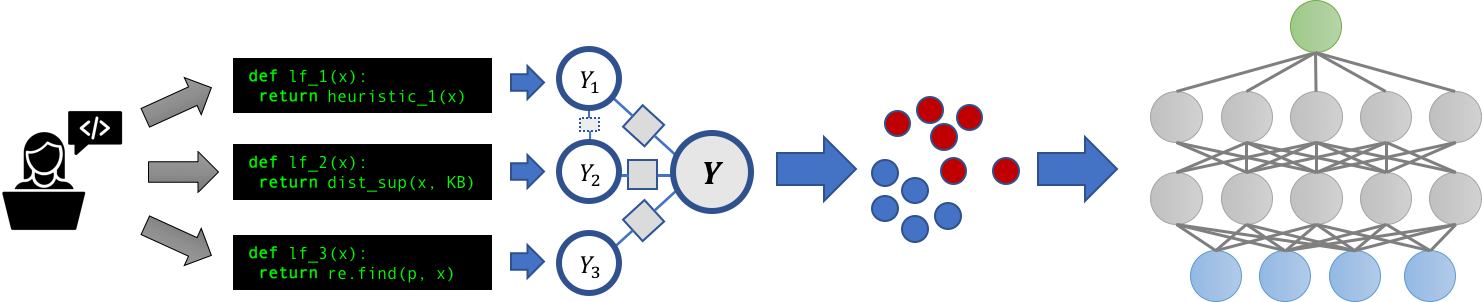
Figure 2: Description of the snorkel pipeline. (1) Users write a set of labeling functions to noisily label their data. (2) A discriminative model is trained based on these labeling functions to produce probabilistic labels for a training set. (3) An end model is trained on these probabilistic labels to be the final classifier
Project Objectives
- Understand how natural language processing models can be used in environments with relatively low annotated data
- Understand what linguistic features are important to gain signal from an unlabeled corpus
- Do word embeddings help give better signal than looking for keywords?
- Do contextual embeddings give better signal than word embeddings, since sequence classification is heavily dependend on the context that the sequence appears in?
- Do POS tags, constituencey parses, and dependency parses provide structural information that gives better signal than just semantic information in the word embedding space?
- Understand how to best generate labeling functions from a small set of annotated instances
Minimum Viable Action Plan
- Build a supervised pipeline for NER and other sequence classification tasks with BIO encoding
- Evaluate a benchmark with random sampling to see how this compares against dataset sizes in
{1 ... 1000}, with a heavier focus on sizes between{1 ... 100} - Investigate semi/weak supervision (presence of a small annotation set and a noisy set), with the following labeling functions
- Every positively annotated word gets stored in a dictionary and is in the noisy set is assumed to be true
- The labeling function labels words that have already been annotated as positive
- Expand the dictionary above with word embeddings (GLOVE) to hopefully gather more data
- The labeling functions labels words/phrases that are close to already labeled words in the word embedding space
- Expand the dictionary above with contextual word embeddings (ELMo/BERT) to hopefully be more relevant in context
- The labeling function labels words/phrases that appear in a similar context to the already labeled words, where similar context is measured by euclidean distance or cosine similarity
- Investigate some structural method (described in stretch goals)
- Every positively annotated word gets stored in a dictionary and is in the noisy set is assumed to be true
Note: for each of the described methods above, in dictionary classifiers and structural methods, we will represent each of these as a labeling function and investigate various combinations of the labeling functions
Stretch Goals
Assuming the minimum viable plan goes as expected, the stretch goals are the following
- Look into POS tags and see if patterns can be drawn there
- Labeling functions could check to see if the POS tags match a regex patterns (e.g.
NN*would refer to any noun phrases).
- Labeling functions could check to see if the POS tags match a regex patterns (e.g.
- Look into constituency and dependency parses
- Labeling functions could rely on Tree Kernels
- Look into a mix of structural and word embedding based methods
Methodologies
Implementation: We will build our models in PyTorch [12 PyTorch] and using AllenNLP [10 Gardner et al. 2018]. We will build our system ontop of Snorkel [1 Ratner et al. 2017]. We will compare our implementation against Snuba/Reef [2 Varma et al. 2018] and AutoNER [9 Shang et al. 2018], and use Snorkel MeTal [11 Hancock et al. 2019] (an implementation of Snorkel for GLUE) to help with the finer grained details of writing models for sequence classification.
Resources
Training Data: We will evaluate our pipelines on the, CADEC [7 Karimi et al. 2015] Adverse Drug Reactions tag, and CONLL [8 Sang et al. 2003] PER tag (identifying people).
Computational Resources: The methods we propose do not rely on large amounts of computational resources, single GPU machines or maybe even CPU machines should be sufficient in the proposed project.
Existing Codebases:
- Snorkel [1 Ratner et al. 2017] is a publicily available project (python library) that we intend to use
- SwellShark [3 Alan et al. 2017] is a paper that describes how Snorkel can be applied to sequence classification problems (biomedical NER), and we will follow the pipeline described there
- AutoNER [9 Shang et al. 2018] is released on github, and describes an implementation of a CRF that relies on probabilistic labels (Fuzzy CRF) as compared to gold labels. We will use this implementation in our experiments.
Evaluation
We will evaluate our project through comparing F1 score with amount of annotated training data for a comparison. We will compare our methods against various benchmarks to see how it performs, in particular:
- Supervised learning on the datasets
- Active learning (no noisy set) on the datasets
- AutoNER and SwellShark reported performances
- Snuba/Reef
- Various combinations of labeling functions described in the Minimum Viable Plan
Related work
Similar work has been done in Snuba/Reef [2 Varma et al. 2018] for text classification through bag of word features, however bag of words is not expressive enough for sequence modeling since context of the words and order of them matters. SwellShark [3 Alan et al. 2017] automatically generates a series of labeling functions for Biomedical NER, which is a promising start, however it relies on access to an external knowledge base, a rather limited set of labeling functions, and has been hand tuned for the tasks at hand. We hope to extend this in a more general setting by looking at (1) a more complex DSL by taking into account linguistic features such as (POS, Constituency Parse Trees, Dependency Parses) and (2) no reliance on an external knowledge base (KB). Babble Labble [3 Hancock et al. 2018], describes a process in which users provide natural language explanations for a label, and labeling functions are generated based off of these descriptions. We will use the insights in this paper to see what features they extracted from explanations and see if they can be generalized, however our pipeline itself will not rely on explanations provided by users. Lastly, AutoNER [9 Shang et al. 2018] relies on no human annotation but the presence of external dictionaries. We hope to rely on a small amount of human annotation as comapred to any external sources such as KB and dictionaries.
References
- Snorkel: Rapid Training Data Creation with Weak Supervision
- Alexander J. Ratner and Stephen H. Bach and Henry R. Ehrenberg and Jason Alan Fries and Sen Wu and Christopher R’e
- 2017 VLDB
- Project Website
- Paper
- Snorkel Reef/Snuba (Snuba: Automating Weak Supervision to Label Training Data):
- Paroma Varma and Christopher R’e
- 2018 PVLDB
- Paper
- SwellShark: A Generative Model for Biomedical Named Entity Recognition without Labeled Data
- Jason Alan Fries and Sen Wu and Alexander J. Ratner and Christopher R’e
- 2017 CoRR
- Paper
- Snorkel Labeling Functions Workshop
- Training Classifiers with Natural Language Explanations (Babble Labble Snorkel)
- Braden Hancock and Paroma Varma and Stephanie Wang and Martin Bringmann and Percy S. Liang and Christopher R’e
- 2018 ACL
- Paper
- No Training Required: Exploring Random Encoders for Sentence Classification
- John Wieting and Douwe Kiela
- 2018 CoRR
- Paper
- Cadec: A corpus of adverse drug event annotation
- Sarvnaz Karimi and Alejandro Metke-Jimenez and Madonna Kemp and Chen Wang}
- 2015 Journal of biomedical informatics
- Paper
- CONLL Dataset (Introduction to the CoNLL-2003 Shared Task: Language-Independent Named Entity Recognition)
- AutoNER
- AllenNLP
- Matt Gardner and Joel Grus and Mark Neumann and Oyvind Tafjord and Pradeep Dasigi and Nelson F. Liu and Matthew E. Peters and Michael Schmitz and Luke S.
- 2018 CoRR
- Paper
- Project Website
- Snorkel MeTal
- PyTorch: pytorch.org/